Someone asked me the other day what the relationship was between the steady-state fluxe through a reaction and the coresponding level of enzyme. Someone else suggested that there would be a linear, or proportional relatinship between a flux and the enzyme level. However, this can’t be true, at least at steady. Considder a 10 step linear pathway. At steady-state each step in the pathway will, by defintion, carry the same flux. This is true even if each step has a different enzyme level. Hence the relationship is so simple. In fact the flux a given step carries is a systemic properties, dependent on all steps in the pathway. As an experiment I decided to do a simulation on some synthetic netowrks with random parameters and enzyme levels. For this exmaple I just used a simple rate law of the form: $$ v = e_i (k_1 A - k_2 B) $$ For a bibi reaction, A + B -> C + D, the coresponding rate law would be: $$ v = e_i (k_1 A B - k_2 C D) $$ a similar picture would be seen for the unibi and biuni reactions. Using our teUtils package I generated random networks with 60 species and 150 reactions. The reactions allowed are uiui-uinbi, biui or bibi. I then randomized the values for the enzymne levels $e_i$ and computed the steady-state flux. I used the following code to do the analysis. I have a small loop that generates 5 random models but obviously this number can be changed. I generate a random model, load the model into roadrunner, randomize the values for the $e_i$ parameters between 0 and 10, compute the steady-state (I do a presimulation to help things along) and collect the corresponding $e_i$ and flux values. Finally I plot each pair in a scatter plot.
import tellurium as te
import roadrunner
import teUtils as tu
import matplotlib.pyplot as plt
import random
for i in range (5):
try:
J = []; E = []
antStr = tu.buildNetworks.getRandomNetwork(60, 150, isReversible=True)
r = te.loada(antStr)
n = r.getNumReactions()
for i in range (n):
r.setValue ('E' + str (i), random.random()*10)
m = r.simulate(0, 200, 300)
r.steadyState()
for i in range (n):
J.append (abs (r.getValue ('J' + str (i))))
E.append (r.getValue ('E' + str (i)))
plt.figure(figsize=(12, 8))
plt.plot(E, J, '.')
except:
print ('Error: bad model')
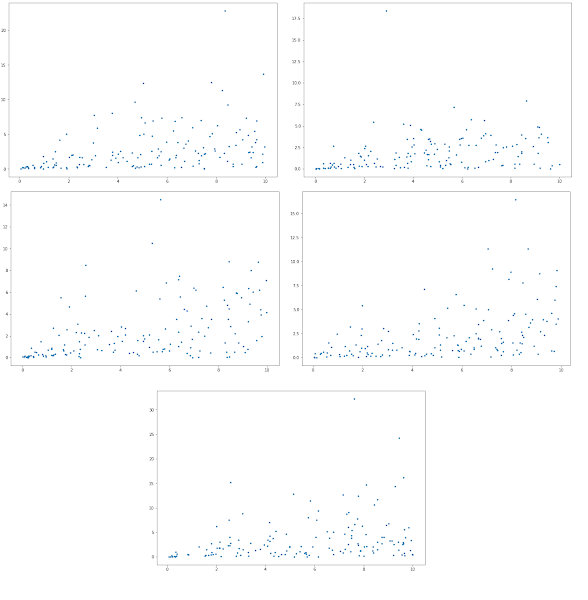
The results for five random networks is shown below. Note the x axis is the enzyme level and the y axis the corresponding steady-state flux through that enzyme. It's intersting to see that there is a rough correlation between enzyme amount and the corresponding flux, but its not very strong. Many of the points are just scattered randomly with some showing a definite correlation. The short answer is the realtinship is not so simple.